Protons are widely used in cancer treatment because of their advantageous interaction properties compared to photons in conventional radiotherapy (Wilson 1946, Newhauser and Zhang 2015). Nevertheless, the exploitation of the full potential offered by protons in clinical practice is limited by several sources of treatment uncertainties (Knopf and Lomax 2013, Parodi 2016, Parodi and Polf 2018, Lomax 2020). Range verification and dose monitoring in proton therapy are thus of interest to fully exploit the favorable properties of proton beams.
Since protons stop inside the patient, it is not possible to detect them directly for range verification purposes. Among different proposed approaches, one promising possibility for monitoring is through secondary prompt gammas (PGs) (Pinto 2024). Range verification with PG was first proposed by Stichelbaut and Jongen (2003), then experimentally verified by Min et al (2006), and subsequently investigated by many groups using instrumentation able to detect different types of signatures, such as spatial distribution just as examples, Testa et al 2008, Richter et al 2016, Xie et al 2017, time (Golnik et al 2014) and energy (Hueso-Gonzalez et al 2018). In this work, the spatial distribution is considered: the emission of PG is correlated with the penetration path of protons in tissue, and the measurement of PG can be used to draw conclusions on the proton range.
PG emission happens in a timescale below the nanosecond. This makes PG suitable for the final purpose of real-time adaptive particle therapy applications, towards which the radiotherapy community interest is increasing. Building upon the benefits seen from online adaptive proton therapy (Albertini et al 2020, Paganetti et al 2021), real-time adaptive particle therapy aims at reducing proton therapy uncertainties not only among treatment fractions, but during a single fraction itself, on a pencil-beam delivery scale. This work is thought to represent a step forward this direction.
For range verification purposes, the expected PG signals related to the dose distributions calculated by the treatment planning program are needed, in order to compare the measurements results with the expected ones and check for mismatches. This approach can be seen as an indirect method for treatment verification. The modeling of the expected PG distributions could be performed by a full Monte-Carlo (MC) simulation. However, this way of proceeding is subject to approximation issues inherent in MC models (Verburg et al 2012). In fact, Verburg et al (2012) showed for example that the nuclear reaction models built in different MC tools predict different PG emissions. Moreover, a full MC calculation of highest accuracy is time consuming, so not suitable for the final goal of real-time verification. To overcome MC limitations, deep learning approaches were proposed (Hu et al 2020, Liu and Huang 2020, Zhang et al 2024), which nevertheless are still in an early stage of research and suffer nowadays from patient specificity and lack of available data for training. An alternative way of proceeding, which could also aid in the training of deep learning approaches, is by means of a filtering approach. The formalism was first introduced by Parodi and Bortfeld (2006) to relate the positron emission tomography (PET) activity profiles to the depth-dose profile delivered during proton irradiation. The main idea behind the filtering is the possibility to determine a PET profile from a dose distribution by performing a 1D convolution with a so-called filter function f. Validation of this approach and extension to PG came with successive works (Attanasi et al 2011, Frey et al 2014, Hofmann et al 2019a, Pinto et al 2020).
PG distributions can then give a direct information about the range of the primary protons. For this reason, the filtering can be mathematically referred to as direct problem. However, other than range, one of the main quantities of interest for cancer therapy is the deposited dose. Getting a direct information on the dose deposition from the PG emission is not as straightforward: the underlying mechanisms of these two physical phenomena are different, as they are governed by nuclear and electronic interactions, respectively. In principle, it would be possible to exploit the introduced filtering formalism inversely, setting up in this way a mathematical inverse problem, using an inverse filter function f−1 to obtain the actual delivered dose. However, this operation is a deconvolution problem, which is ill-posed: there are infinite possible solutions, sensitive to noise in particular.
To face this challenge, several deconvolution techniques have been proposed for different secondary emission signals. In particular, the following three approaches were identified in this work as most promising for PG-based dose reconstruction (i.e. reconstruction of dose distribution): the analytical deconvolution method (Remmele et al 2011), the evolutionary algorithm (Schumann et al 2016, Hofmann et al 2019b, Yao et al 2023) and the maximum-likelihood expectation-maximization (MLEM) algorithm (Masuda et al 2019, 2020), which were applied in literature to reconstruct the dose delivered by ions from secondary radiation data, mostly PET. They can all be built around the filtering formalism, which is then crucial for their implementation, and they can also potentially be integrated in a treatment planning system, as it already happened for the filtering (Pinto et al 2020). The analytical deconvolution and the MLEM algorithm were applied to reconstruct the dose delivered by protons from PET signals, generated through simulations and from measurements in homogeneous and heterogeneous phantom (Remmele et al 2011, Masuda et al 2019), from head and neck (H&N) phantom simulations for the case of MLEM (Masuda et al 2020), and from measurements in the case of a computed tomography (CT) data of a H&N patient for the deconvolution approach (Remmele et al 2011). The evolutionary algorithm was first introduced to predict the depth-dose profile from simulated PG distributions in water phantoms (Schumann et al 2016), then the same approach was also explored to reconstruct the dose profile from PET measurements generated after interactions of carbon ions (Hofmann et al 2019b). More recently, a double evolutionary algorithm was used to realize a PG-based reconstruction of simulated dose in a thoracic CT considering a non-ideal Compton camera (Yao et al 2023).
In the present work, all the abovementioned algorithms are first implemented and applied to simulated dose deposited in homogeneous and inhomogeneous phantoms, for pristine Bragg peaks and spread-out Bragg peaks (SOBPs). Both laterally integrated 1D dose-depth profiles and 3D dose distributions were reconstructed. To investigate the 3D applicability of the dose reconstruction methods, in silico data were also generated on the CT data of a H&N patient. The performance of the techniques is evaluated and compared in terms of different metrics. Before presenting the obtained results in section 3, a brief recall of the established filtering approach is given in section 2, followed by the description of how the different dose reconstruction approaches considered herein work. Section 4 compares the methods and provides comments about their potential, giving an outlook on future steps.
2.1. Direct problem: filteringThis section focuses on the forward generation of depth-PG distributions from depth-dose profiles, similar to the work by Schumann et al (2016) and Pinto et al (2020). Here, the main ideas of the forward filtering framework are described, as it is an important part of the dose reconstruction workflows described in the following sections. For a more detailed description of the mathematical basics of the approach, please refer to Parodi and Bortfeld (2006) and successive works (Attanasi et al 2011, Frey et al 2014, Hofmann et al 2019a, Pinto et al 2020).
First, a proton beam irradiating a homogeneous target is taken into account. The 1D PG signal P(z) along the beam propagation depth z can be obtained from the corresponding 1D dose distribution D(z) by convolving (i.e. filtering) the dose profile itself with a filter function f(z):

To estimate this filter function, in principle it would be possible to start from simulated data and then proceed with deconvolution methods. However, this is an ill-posed problem. The alternative is to move on analytically. The key elements of this analytical formalism are the so-called 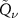 functions, defined as the convolution of Gaussian G(x) and powerlaw functions
functions, defined as the convolution of Gaussian G(x) and powerlaw functions 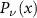 :
:

The parameter ν is called shape and defines the peak-to-plateau ratio of the profile. Shifted and scaled versions of these 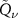 functions are first used to fit the simulated depth-dose and depth-PG profiles. A
functions are first used to fit the simulated depth-dose and depth-PG profiles. A 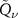 function can then be described with four parameters: an overall normalization factor or weight w, the shape ν, the shift a as measure of the horizontal shift of the depth profiles, and the scale σ, which defines the width of the distal peak.
function can then be described with four parameters: an overall normalization factor or weight w, the shape ν, the shift a as measure of the horizontal shift of the depth profiles, and the scale σ, which defines the width of the distal peak.
Then, a useful mathematical property of the 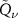 functions is exploited: in fact, the convolution of two
functions is exploited: in fact, the convolution of two 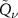 functions is still a
functions is still a 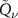 function, so
function, so
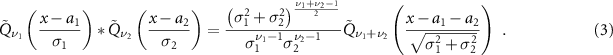
That said, if P(z) and D(z) are expressed as 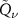 functions, then f(z) will be a
functions, then f(z) will be a 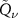 function as well. The
function as well. The 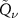 function’s parameters of the filter can then be found exploiting the following relationships: if P indicates quantities related to the PG distribution, D to the dose profile and f to the filter,
function’s parameters of the filter can then be found exploiting the following relationships: if P indicates quantities related to the PG distribution, D to the dose profile and f to the filter,

These equations imply the following conditions:

One filter must be considered for each kind of nucleus in the target which emits PG after a nuclear interaction induced by a proton. The filters are also dependent on the energy of the primary protons. This has been previously showed by Schumann et al (2016), and mainly the parameters w and a are concerned. To tackle this filter energy dependence, the filter functions were here optimized for energies spanning the treatment energy window, from 110 MeV to 230 MeV. This was sufficient for the purposes of the current work, as showed in section 3.1.
The extension of the filtering to different homogeneous targets than the one considered to compute the filters and to inhomogeneous cases is straightforward (Parodi and Bortfeld 2006), and it can be performed still exploiting the filter functions computed for one homogeneous material, with additional steps. First, a water-equivalent range-conversion formalism is necessary. Protons irradiating different tissues with the same initial energy stop at different positions along the beam direction. Knowing the path length of protons with a defined initial energy in a certain material, it is possible to obtain the corresponding path length l traversed by protons in a second material taken as reference (usually water) as

where 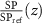 is the relative stopping power between the traversed material and the reference one at the depth z′ of the traversed medium. The filtering can then be applied in the homogeneous, reference space. Dose needs to be rescaled when moving from the target space to the reference one, exploiting the relative mass stopping power. To analytically calculate PG in actual space, one eventually has to consider the weight for each nucleus relative to the tissue composition of the original, reference homogeneous material (for which the filters were calculated). This weight can be introduced by a local factor
is the relative stopping power between the traversed material and the reference one at the depth z′ of the traversed medium. The filtering can then be applied in the homogeneous, reference space. Dose needs to be rescaled when moving from the target space to the reference one, exploiting the relative mass stopping power. To analytically calculate PG in actual space, one eventually has to consider the weight for each nucleus relative to the tissue composition of the original, reference homogeneous material (for which the filters were calculated). This weight can be introduced by a local factor 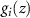 :
:

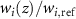 is the weight fraction of the target nucleus involved in the reaction channel in the actual material at depth z relative to the reference homogeneous material, while
is the weight fraction of the target nucleus involved in the reaction channel in the actual material at depth z relative to the reference homogeneous material, while 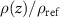 is the mass density of the material at depth z relative to to the reference one.
is the mass density of the material at depth z relative to to the reference one.
To take into account the presence of heterogeneities in all directions, and not just along the beam direction, a raytracing operation is performed. This also makes possible to analytically calculate 3D PG distributions from 3D dose distributions. By raytracing, the proton beam is divided into N narrow sub-beams, or rays, each representing a straight line in the target. The range conversion and the gi factors are applied to each ray, which are then traced from the actual to the reference, homogeneous space. Summing up all the sub-beam doses in reference space, the laterally integrated depth-dose profile in reference space can be obtained. The laterally integrated depth-PG profiles for each nucleus of the reference target are then calculated via filtering. Eventually, it is possible to obtain the analytical 3D PG distribution from the laterally integrated depth-PG profile analytically calculated tracing every sub-beam back in the actual space by inversely applying the range conversion and gi factors to each of them.
2.2. Inverse problem: dose reconstructionWith the described filtering approach, the PG signal at emission can be estimated from the dose distribution and indirect range verification operations can be carried out by comparing the said PG curve with a possible measured one, corrected for the detector response. This comparison can then give information on the difference between the measured range of protons and the expected one, but no knowledge would be available regarding the dose itself. In order to reconstruct the dose from a PG distribution via the filtering formalism, several alternative methods have been proposed. This section focuses on the description of the three methods, considered herein as most promising because of their relationship with the described filtering formalism. They exploit the application of a suitable deconvolution algorithm, which is an inverse problem and, after its application, noise or other artifacts in the data may be amplified, leading to a degraded reconstruction. This is why a regularization term is introduced, which uses a prior information on the dose distribution. Before describing the individual reconstruction methods, an introduction of regularization as common to all the considered algorithms is provided.
2.2.1. RegularizationA regularization term is included in all reconstruction algorithms, which implies the consideration of a prior knowledge of the signal, in order to stabilize the problem by adding some constraints to restrict the amount of the possible solutions.
When it is reasonable to assume a smooth solution, as for the case of dose profiles, the Tikhonov–Miller (TM) (Press et al 2002) and the total-variation (TV) regularizations (Vogel 2002) are suitable for noise amplification suppression in deconvolution problems. Both TM and TV use the gradient 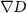 of the reconstructed dose signal D to find the a priori knowledge. The TM regularization is expressed as a L2 norm:
of the reconstructed dose signal D to find the a priori knowledge. The TM regularization is expressed as a L2 norm:

while the TV regularization is a L1 norm:

The TM regularization has the effect of smoothing edges. The TV regularization was proposed to overcome this problem, but once applied, a staircasing effect occurs in the case of a continuous increase of signal values. That said, the choice of the regularization term depends on the features of the given signals. The TV regularization should be used in the case of sharp edges, while the TM regularization is recommended in the case of a continuous increase of signal values. For these reasons, and since its use was well motivated in the previous literature for the deconvolution approach (Remmele et al 2011) and the evolutionary algorithm (Hofmann et al 2019b) described in the following sections, the TM regularization was chosen for the implementation of the dose reconstruction algorithms explored and described in the following. TV regularization was also described for a matter of completeness.
2.2.2. Analytical deconvolution approachThe analytical deconvolution approach, initially developed by Remmele et al (2011) in the context of PET verification, consists of an iterative maximum a posteriori approach based on the minimization of the functional J, defined as:

R is the TM regularization term, which is balanced against S by a parameter λ. A good value for λ can be found via a ‘brute-force’ search. S is called similarity (or fitting) term, and realizes the comparison between the expected PG signal, coming from the filtering of the dose to be reconstructed, and the simulated (potentially measured) one. The definition of S depends on the underlying noise distribution. Since the PG detection is a photon detection technique involving a low number of counts, measured signals are influenced by Poisson noise, and Gaussian noise may be associated with the detector’s performance. If the Poisson noise outweighs the Gaussian one, as it can be for acquired signals, then the similarity term is better expressed as a Richardson–Lucy term:
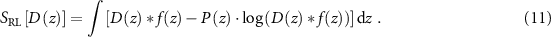
Within this work, high-statistics simulated data were considered, where the Gaussian noise overcomes the Poisson one. A more suitable definition of S in these conditions is accomplished with a least-squares (LSs) term:
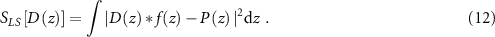
Remmele et al (2011) pointed out that the noise amplitude in the reconstructed dose reflects the noise in PET signals, and it is not possible to completely eliminate it. The same concept remains valid for PG signals. Higher noises can be smoothed with a higher regularization, which means a higher value of λ, but a perfect solution is not achievable. In this work, an attempt in trying to face this issue was made exploiting the 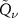 functions formalism. The dose distribution was first described using one
functions formalism. The dose distribution was first described using one 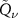 function. Then, instead of optimizing the similarity metrics bin by bin in the dose distribution as in Remmele et al (2011), the optimization was carried out in terms of
function. Then, instead of optimizing the similarity metrics bin by bin in the dose distribution as in Remmele et al (2011), the optimization was carried out in terms of 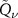 function’s parameters. This process is expected to be faster, because the optimization involves only the four
function’s parameters. This process is expected to be faster, because the optimization involves only the four 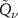 function’s parameters instead of all bins in depth. Another advantage is that noise is not reflected in the reconstruction. However, what has to be kept in mind is that a
function’s parameters instead of all bins in depth. Another advantage is that noise is not reflected in the reconstruction. However, what has to be kept in mind is that a 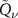 function is already an approximation itself, so even in this case a perfect match is not achievable. Results related to this new approach and a comparison with the more standard, established one from Remmele et al (2011) are showed in section 3.2.
function is already an approximation itself, so even in this case a perfect match is not achievable. Results related to this new approach and a comparison with the more standard, established one from Remmele et al (2011) are showed in section 3.2.
The analytical deconvolution approach relies on an optimization process. To implement it, the minimize function within the scipy.optimize Python package was used. The Nelder–Mead optimization algorithm was chosen. The standard deconvolution algorithm from Remmele et al (2011) was compared with the new strategy of deconvolution performed only in terms of 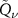 function parameters. For this latter algorithm, there is no need of regularization. The Nelder–Mead optimization algorithm was used also in this case. To define convergence of these two deconvolution algorithms, the tolerance value, which sets the acceptable error on both the optimized variable (i.e. D(z)) and the minimized cost function (i.e.
function parameters. For this latter algorithm, there is no need of regularization. The Nelder–Mead optimization algorithm was used also in this case. To define convergence of these two deconvolution algorithms, the tolerance value, which sets the acceptable error on both the optimized variable (i.e. D(z)) and the minimized cost function (i.e. ![$J[D(z)]$](https://content.cld.iop.org/journals/0031-9155/70/23/235026/revision2/pmbae1fcbieqn26.gif) ), was set at 10−6.
), was set at 10−6.
The evolutionary algorithm can be considered as a gradient-free optimization approach as well, as it relies on an iterative process. The algorithm begins with the generation of an initial population of N individuals, each of them representing one depth-dose profile 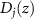 . These initial profiles are first randomly mutated. Mutations can be of any nature. In this work, as in Hofmann et al (2019b), only one of the following mutation at a time was applied: a random horizontal shift in depth of integer bins in a range from −2 to 2; a random vertical scale of a factor between 0.8 and 1.2; or a local variation, which consists in adding a Gaussian distribution to a randomly chosen point, with random height (up to
. These initial profiles are first randomly mutated. Mutations can be of any nature. In this work, as in Hofmann et al (2019b), only one of the following mutation at a time was applied: a random horizontal shift in depth of integer bins in a range from −2 to 2; a random vertical scale of a factor between 0.8 and 1.2; or a local variation, which consists in adding a Gaussian distribution to a randomly chosen point, with random height (up to 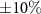 of the dose at the chosen bin) and σ within 10 sampled bins. The mutated dose individuals of the population are then convolved with the filter functions, in order to obtain a first population of predicted depth-PG profiles
of the dose at the chosen bin) and σ within 10 sampled bins. The mutated dose individuals of the population are then convolved with the filter functions, in order to obtain a first population of predicted depth-PG profiles 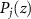 . Each of these jth depth-PG predictions is compared to the ground-truth one, potentially measured. The agreement of each comparison j at every iteration k is assessed by a quantity called fitness value
. Each of these jth depth-PG predictions is compared to the ground-truth one, potentially measured. The agreement of each comparison j at every iteration k is assessed by a quantity called fitness value 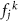 , defined as
, defined as
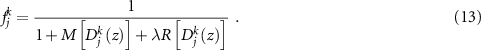
M can be any metric used to compare two curves. In this work, the normalized root mean squared error (NRMSE) was used. R and λ are the regularization term and its weighting factor, respectively, analogous to what was introduced in section 2.2.2. The fitness value is a criterion which quantifies the quality of the reconstructed dose for that individual by comparing the depth-PG profile obtained after the filtering of that reconstructed dose and the ground-truth depth-PG profile. The higher the fitness of an individual of the population is, the more similar is the ground-truth depth-PG profile to the one calculated from filtering of that same individual. To then select which individuals will be the parents for the following iteration, a fitness selection is applied. These new parents individuals are then randomly mutated again, in order to vary their profile and reach a potential improvement in fitness. These steps are repeated until a certain termination condition is reached, which could be related to the fitness value or to the number of iterations. Eventually, the individual with the higher fitness in the population is designated as the reconstructed dose.
For the evolutionary algorithm, a population of N = 300 individuals was considered. Increasing the number of individuals in a population improves the fitness, but also increases the computing time linearly. As in Hofmann et al (2019b), the probability for the occuring mutations was set at 0.3, 0.3 and 0.4 for the horizontal shift, the scaling and the local Gaussian variation, respectively. The algorithm was stopped after 1500 iterations and, due to its stochastic nature, it was repeated 10 times, with different initial conditions introduced in the first random mutation. A single result among the 10 was eventually chosen, using the criterion of the lowest NRMSE between the laterally integrated profiles of the simulated PG and the PG calculated after filtering of the dose at the last iteration. The initial parameters were at first chosen from the literature (Hofmann et al 2019b) and then varied empirically (between 100–500 individuals and 500–3000 iterations) until a good compromise between computational effort and quality of the result was achieved.
2.2.4. MLEM algorithmThe MLEM algorithm is better known for its clinical use in image reconstruction. Masuda et al (2019) gave it a different purpose, exploiting it for dose reconstruction. The inherent filtering is here implemented with a different approach. The convolution in its common mathematical expression is not performed anymore. Instead, the equivalent matrix-vector product of the dose profile with a system matrix C is considered. If we define δj the dose in a certain voxel j and pij its corresponding PG yield, then pij can be calculated as

If Pi is the actual measured PG signal in the voxel i, the expected PG signal 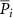 coming from filtering can be computed as
coming from filtering can be computed as

The construction of C is based on the generated filter function. When more tissues and inhomogeneous targets are taken into account, the range conversion and local factor application described in section 2.1 are performed in the system matrix itself (Masuda et al 2020).
The dose reconstruction algorithm is based on the maximization of the likelihood, defined as the probability that a certain PG distribution 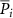 obtained from applying the filtering to a dose distribution corresponds to the related measured signal Pi. The maximization of this quantity brings to the MLEM formula:
obtained from applying the filtering to a dose distribution corresponds to the related measured signal Pi. The maximization of this quantity brings to the MLEM formula:

Since this is still an inverse reconstruction problem, there is the need to introduce a regularization term, as in section 2.2.1. Masuda et al (2020) applied a TV superiorization to the dose signal itself at each iteration, dividing the MLEM algorithm in two steps and implementing what they referred to as EM-TV algorithm. In this work, the regularization was implemented in a different way. First, a TM regularization was applied as in the other methods, in order to make a meaningful comparison among them. For the same reason, instead of working on the signal itself, the regularization was applied to the objective function, i.e. the likelihood, by implementing a Bayesian reconstruction. To carry this out, the one-step-late approximation, introduced first by Green (1990), was used. This requires the following change in the MLEM formula:
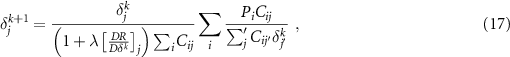
where R is the regularization term, and it is derived with respect to the dose at the previous iteration k.
Before applying the MLEM, the generation of system matrices was necessary. In fact, the MLEM algorithm application foresees that each filter is incorporated in a respective system matrix, whose rows in this case were related to the PG space, and the columns to the dose space. The generation of the system matrices for 1D dose reconstruction is described in appendix A of the supplementary material. Once the system matrices were available, the MLEM algorithm ran for a total of 500 iterations, also determined empirically starting from literature as for the parameters of the evolutionary algorithm, and the selected result was the one at the iteration with lower NRMSE between the simulated PG profile and the PG calculated after filtering of the dose.
2.2.5. Consideration of lateral heterogeneities and 3D reconstructionAs for the forward filtering, each dose reconstruction algorithm can be also applied in 3D, and also when heterogeneities are present in all directions of the target, and not just along the beam direction. The complete workflow to reconstruct 3D dose distributions from 3D PG distributions works as follows:
(i)
An initial guess of the dose (e.g. the treatment plan dose) is considered, and a raytracing operation is performed, to divide the proton beam into N sub-beams (or rays);
(ii)
With the application of range conversion and gi factors, the initial dose of each single ray is converted from actual to reference space;
(iii)
All ray doses in reference space are summed up, and the laterally integrated depth-dose profile in reference space is obtained;
(iv)
Filtering is then applied, which means that the laterally integrated depth-PG profiles for each nucleus of the reference target are analytically calculated;
(v)
The total laterally integrated depth-PG profile in real space is calculated analytically after tracing every sub-beam back by inversely applying the range conversion and gi factors to each ray;
(vi)
With one of the described algorithms, the laterally integrated 1D depth-dose profiles in reference space are then reconstructed;
(vii)
After the 1D reconstruction of the dose in reference space is completed, the inverse application of range conversion and tissue composition information allows to obtain the analytical 3D dose distribution in real space.
It is worth noticing that, while the dose is reconstructed in reference space (as described in point vi) and converted in real space only at the end of the reconstruction (point vii), the iterative comparison of the laterally integrated depth-PG signals inherent in each algorithm happens in real space (point v). The need of moving from one space to another at each iteration for comparing the PG signals is because it is not possible, after potential detection, to distinguish which nucleus a single PG comes from. Moreover, the true relative stopping power and gi factors, which would be needed to convert the PG signals from real to reference space, are not known. With the MLEM algorithm, the laterally integrated 1D depth-dose profiles are reconstructed directly in the actual space, since the information on the range conversion and tissue composition are inherent in the system matrix. When reconstructing with MLEM in 3D, one system matrix is considered for each ray involved in the raytracing process. The quantities necessary to perform raytracing are interpolated on a common array.
A scheme of the 3D reconstruction workflow is shown in figure B1 provided in appendix B of the supplementary material.
2.3. Voxelized phantoms simulationsThe simulated dose and PG distributions in phantoms were obtained with the general purpose MC toolkit Geant4 (Agostinelli et al 2003), version 10.07.p03, using the physics list QGSP_BIC.
First, a rectangular phantom of 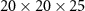 cm3 in x, y and z, respectively, made of an homogeneous reference material (referred to as REFMAT in this paper) was created. The properties of REFMAT are listed in table 1. The choice of this particular REFMAT is the same as in Hofmann et al (2019a) and Pinto et al (2020) and it takes into consideration the six most abundant elements in tissues, which produce a consistent amount of PG.
cm3 in x, y and z, respectively, made of an homogeneous reference material (referred to as REFMAT in this paper) was created. The properties of REFMAT are listed in table 1. The choice of this particular REFMAT is the same as in Hofmann et al (2019a) and Pinto et al (2020) and it takes into consideration the six most abundant elements in tissues, which produce a consistent amount of PG.
Table 1. REFMAT properties. Density = 1.54 g cm−3.
ElementFraction by mass (%)1 H1.7616C30.814N25.916O37.731P1.3440Ca2.5Then, two inhomogeneous phantoms were generated. Phantom1 has only longitudinal inhomogeneities. It is made of six slabs made of soft tissue, bone, lung tissue, bone and soft tissue. Materials were chosen from the Geant4 libraries, in particular ‘G4_TISSUE_SOFT_ICRP’, ‘G4_BONE_CORTICAL_ICRP’ and ‘G4_LUNG_ICRP’. Phantom2 was inspired by Parodi et al (2005). It is composed of slabs disposed in a way to generate lateral heterogeneities. It is made of polyethylene (PE, ‘G4_POLYETHYLENE’), polymethylmethacrylate (PMMA), muscle tissue (‘G4_MUSCLE_SKELETAL_ICRP’), bone tissue and lung tissue. The geometry of Phantom1 and Phantom2 can be visualized with the results in section 3.1.
The simulations consider a proton pencil beam shot onto the target xy front face, along the z-axis. The total volume of all three targets was divided in 1 mm3 cubic sub-volumes, building a voxelized structure. The energy deposition, consequently the dose, and the information for the emitted photons, like the coordinates at emission or the nucleus involved in the reaction, were scored for each voxel. Only photons created after hadronic interactions of protons with target nuclei were considered in the analysis. Depth-dose and PG profiles were created by means of lateral integration. Emitted PG were scored within a certain energy window only, between 1 MeV and 10 MeV, in order to consider for most common PG monitoring techniques responses (Pinto et al 2020 and references therein). This energy window can be modified as necessary by exploiting the look-up table formalism developed by Pinto et al (2020). This allows for a swift application of the approaches presented herein to the different PG cameras and energy windows considered in PG monitoring by different authors (just as examples, Richter et al 2016, Xie et al 2017, Hueso-Gonzalez et al 2018, Missaglia et al 2023).
2.4. Filter creation and scenarios for dose reconstruction in the voxelized phantomsThe filter functions, one for each target nucleus present in REFMAT (except hydrogen), were created by irradiating the REFMAT with 109 protons of incident energy equal to 150 MeV. The filters creation process is mentioned in section 3.1 and explained more in detail in appendix A of the supplementary material. To tackle the energy dependence of the filters, they were optimized considering four energies in the therapeutic energy window. So, protons incident on REFMAT with energies equal to 110 MeV, 190 MeV and 230 MeV other than 150 MeV were also simulated, and the related depth-PG and dose profiles were used for energy optimization of the filters. This optimization was finally tested for 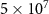 protons impinging on a homogeneous phantom of REFMAT with an energy of 200 MeV.
protons impinging on a homogeneous phantom of REFMAT with an energy of 200 MeV.
All the dose reconstruction approaches were first applied to REFMAT and to Phantom1. A first scenario foresees the reconstruction methods applied to a single pristine Bragg peak, made of a beam comprising 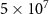 particles incident perpendicularly to the xy surface of the phantoms with an energy of 200 MeV and a 2D Gaussian shape with a σ of 4 mm. To check for the sensitivity of the methods to small range variations, range shifts were approximated by a change in the initial energy of the proton beams, to represent a global change. Protons incident on REFMAT with energies from 198.5 MeV to 201.5 MeV with a step of 0.5 MeV were then simulated, corresponding to expected range shifts in water of about 1 mm. The range shifts were evaluated as
particles incident perpendicularly to the xy surface of the phantoms with an energy of 200 MeV and a 2D Gaussian shape with a σ of 4 mm. To check for the sensitivity of the methods to small range variations, range shifts were approximated by a change in the initial energy of the proton beams, to represent a global change. Protons incident on REFMAT with energies from 198.5 MeV to 201.5 MeV with a step of 0.5 MeV were then simulated, corresponding to expected range shifts in water of about 1 mm. The range shifts were evaluated as 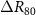 and
and 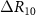 , with
, with 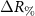 being the difference of the positions at the % of the dose maximum in the distal fall-off region between the expected curve at 200 MeV and the ones reconstructed from the PG distributions at the actual energies. A positive value indicates a higher range for the expected curve than the reconstructed one. The real range shifts relative to the planned energy were also obtained from simulated data and were used for comparison.
being the difference of the positions at the % of the dose maximum in the distal fall-off region between the expected curve at 200 MeV and the ones reconstructed from the PG distributions at the actual energies. A positive value indicates a higher range for the expected curve than the reconstructed one. The real range shifts relative to the planned energy were also obtained from simulated data and were used for comparison.
A second scenario considers the reconstruction of a SOBP, simulated following the approach developed first by Bortfeld (1997), then by Jette and Chen (2011). For the SOBP used in this work, 11 different energies (from 159.5 MeV to 180.9 MeV for the generation of the SOBP in REFMAT, from 180.7 MeV to 205 MeV for the one in the Phantom1) were used as contributions, forming a plateau as wide as the 20% of the maximum proton range and with a maximum of 2 Gy, to resemble the maximum fraction dose in a realistic clinical treatment plan. Dose was reconstructed separately for each contribution, to consider a realistic spot-by-spot delivery.
Phantom2 was exploited to extend the reconstruction methods from 1D to 3D, also tackling the presence of lateral heterogeneities, created by how the single slabs are positioned within the phantom. Additional lateral heterogeneities were also introduced simulating the phantom irradiation with different incident angles. Beams with σ from 2 to 10 mm incident on Phantom2 with energies from 110 to 200 MeV and angles of 0, 15, 30 and 45∘ were simulated to thoroughly test the 3D reconstruction algorithm before the extension to patient data.
2.5. In silico patient dataTo extend the 3D dose reconstruction workflow to in silico patient data, a H&N patient treated at the Heidelberg Ion Beam Therapy Center with scanned proton beams (Pinto et al 2020) was considered. All the spots included in the plan were simulated with Geant4 in order to obtain simulated dose and PG 3D distributions. For the purpose of this work, the considered PG distributions only include PGs generated after interaction with nuclei of carbon, nitrogen, oxygen, phosphorus and calcium, since the filters were generated for these nuclei only. Two spots were chosen for a first testing of the algorithms, one in the layer with the highest energy (132.30 MeV) and one in the layer with the lower energy (106.82 MeV). These two spots where chosen becaus
Comments (0)